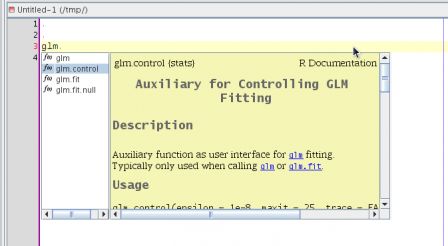
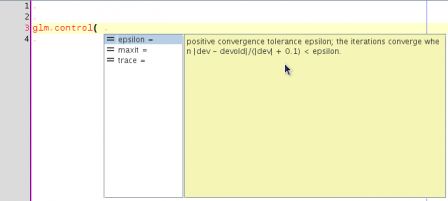
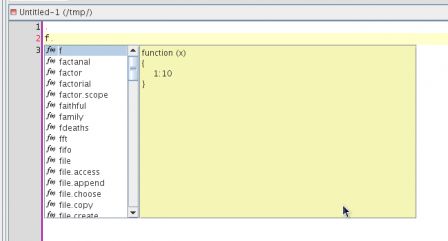
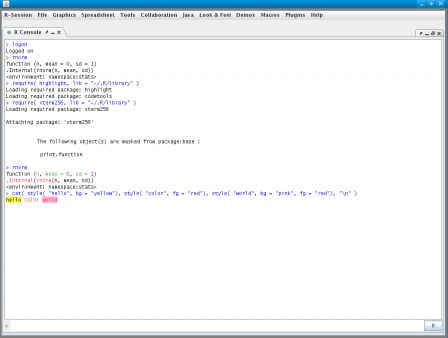
This is a post in the series Mixing R CMD build and ant. Previous posts have shown how to compile the java code that lives in the src directory of a package and how to document this code using javadoc.
This post tackles the problem of unit testing of the java functionality that is shipped as part of an R package. Java has several unit test platforms, we will use junit here, but similar things could be done with other systems such as testng, ...
The helloJavaWorld package now looks like this :
.
|-- DESCRIPTION
|-- NAMESPACE
|-- R
| |-- helloJavaWorld.R
| `-- onLoad.R
|-- inst
| |-- doc
| | |-- helloJavaWorld.Rnw
| | |-- helloJavaWorld.pdf
| | `-- helloJavaWorld.tex
| `-- java
| |-- hellojavaworld-tests.jar
| `-- hellojavaworld.jar
|-- man
| `-- helloJavaWorld.Rd
`-- src
|-- Makevars
|-- build.xml
|-- junit
| `-- HelloJavaWorld_Test.java
|-- lib
| `-- junit-4.7.jar
`-- src
`-- HelloJavaWorld.java
9 directories, 15 files
We have added the src/lib directory that contains the junit library and the HelloJavaWorld_Test.java that contain a simple class with a unit test
And the ant build file has been changed in order to
- build the junit test cases, see the build-testcases target
- run the unit tests, see the test target
- create nice html reports, see the report target
The package can be downloaded here
Coming next, handling of dependencies between java code that lives in different R packages


 Future plans for this package contain:
Future plans for this package contain: